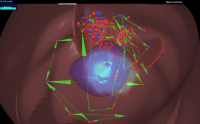
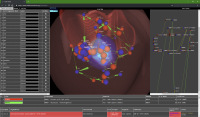
The CELLmicrocosmos 4.2 PathwayIntegration (CmPI) is a tool which provides hybrid-dimensional visualization and analysis of intracellular protein and gene localizations in the context of a virtual 3D environment. This tool is developed based on Java/Java3D/JOGL and provides a standalone application compatible to all relevant operating systems. However, it requires Java and the local installation of the software. Here we present the prototype of an alternative web-based visualization approach, using Three.js and D3.js. In this way it is possible to visualize and explore CmPI-generated localization scenarios including networks mapped to 3D cell components by just providing a URL to a collaboration partner. This publication describes the integration of the different technologies – Three.js, D3.js and PHP – as well as an application case: a localization scenario of the citrate cycle. The CmPI web viewer is available at: http://CmPIweb.CELLmicrocosmos.org.

 JIB Publications
JIB Publications
- Kovanci G, Ghaffar M, Sommer B. Web-based hybrid-dimensional Visualization and Exploration of Cytological Localization Scenarios. J Integr Bioinform. 2016;13(4):47–58. doi 10.2390/biecoll-jib-2016-298; PubMed 28187414
- Sommer B. The CELLmicrocosmos Tools: A Small History of Java-Based Cell and Membrane Modelling Open Source Software Development.. J Integr Bioinform. 2019;. doi 10.1515/jib-2019-0057; PubMed 31560649
Homepage bio.tools
